The Interplay between Lipid Nanodomains, Protein Clusters and Cell Topography in Cell Signalling
Short description
Signal transduction from the environment to cell interior is a fundamental biological process acting across a membrane. Using superresolution microscopy of the type that was awarded a Nobel Prize in 2014, Ingela Parmryd's group studies how signalling molecules in T cells, a cell type specialized in immune defence, are organized before and after the signalling has been initiated. We study the prevalence of molecular clusters, how the clusters are related to the properties of the membrane and change depending on the cellular environment. To interpret the results, we develop image analysis methods of use also for a wide range of other research areas. Increased knowledge on how signalling is initiated is of importance for therapeutical intervention of the aberrant signalling that is a hallmark of multiple serious diseases, including cancer.
The Functional Organisation of the Plasma Membrane in T cell Signalling
The plasma membrane of eukaryotic cells contains nanodomains, commonly referred to as lipid rafts, which are more ordered than the rest of the plasma membrane. The high order is generally considered equivalent to the tight packing of cholesterol and sphingolipids observed in model membranes. However, we have demonstrated that lipid rafts form when actin filaments are pinned to the plasma membrane via phosphoinositides and when extracellularly exposed receptors are pinned by antibodies (Dinic et al., 2013), suggesting that the mechanism for lipid raft formation is lipid-protein interactions.
We have shown that T cell signalling is initiated upon lipid raft aggregation that can be triggered by cold stress and changes in the plasma membrane lipid composition (Magee et al., 2005; Mahammad et al. 2010) and that the T cell receptor in resting T cells resides in lipid rafts that are brought together upon receptor engagement (Dinic et al., 2015). We are now investigating what triggers the formation of lipid rafts in more detail and how membrane order affects the molecular clustering that accompanies T cell signalling using techniques like super resolution microscopy and fluorescence correlation spectroscopy. The cell studies are accompanied by the development of methods for cluster analysis.
Cell Topography and Plasma Membrane Models
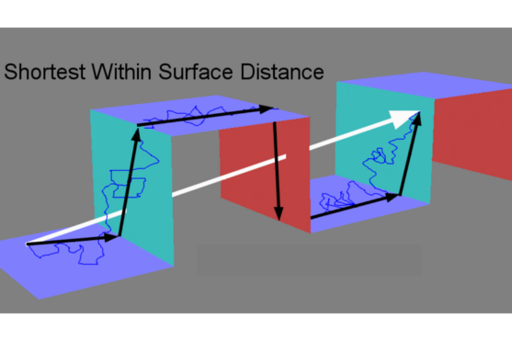
The cell surface is neither flat nor smooth but this is rarely acknowledged. Using high resolution topographical maps of live cells, we and our collaborators have demonstrated that apparent topographical trapping can be behind phenomena described as hop diffusion and transient anchorage in diffusion studies (Adler et al., 2010). Even binding could be the result of apparent topographical trapping when single particle tracks are interpreted in 2D when the molecules are moving over 3D surface. We have shown that ignoring membrane topography causes a consistent underestimation of diffusion and that membrane topography itself can cause anomalous diffusion. Our conclusion is that disentangling apparent from genuine diffusion requires surface characterisation (Adler et al., 2019; Gesper et al., 2020).
Co-localisation Analysis
We develop image analysis methods to get quantitative and objective answers to biological questions. We have developed and patented the method RBNCC (replicate based noise corrected correlation) where image noise, which is unavoidable and leads to the underestimation of any underlying correlation, can be eliminated from correlation measurements (Adler et al. 2008). RBNCC forms the basis for our company No More Noise.
We have performed detailed studies on coefficients developed for use in colocalisation analyses revealing that several are of doubtful value (Adler & Parmryd, 2010; Adler & Parmryd, 2018, Adler & Parmryd, 2019). We were the first to suggest that colocalisation analysis should be divided into the two subgroups co-occurrence and correlation (Adler & Parmryd, 2007; Adler & Parmryd, 2013) and that only pixels where both fluorophores are present should be included in correlation analyses (Adler et al. 2008; Adler & Parmryd, 2014). We now investigate how deconvolution affects correlation analysis and how different intracellular distributions of molecules are represented by our division of colocalisation analysis into co-occurrence and correlation.
Tumour Cell Killing Vδ2Vγ9 T Cells
Vδ2Vγ9 T cells are a T cell subset that recognise and kill cancer cells that accumulate high levels of phosphoantigens, small organic compounds with phosphate groups. There is a positive correlation between the Vδ2Vγ9 T cell number and tumour cell death making Vδ2Vγ9 T cells appealing candidates for immunotherapy. A pilot study indicated that colon cancer patients have lower numbers of circulating Vδ2Vγ9 T cells than healthy individuals.
Together with collaborators at the Uppsala University Hospital we address how the prevalence of Vδ2Vγ9 T cells in colon cancer patients at the four different cancer stages varies and characterise the Vδ2Vγ9 T cells regarding differentiation status, tumour homing potential, proliferation and cytotoxicity.
Together with collaborators at Stockholm University we found that media from erythrocytes infected with P. falciparum can stimulate Vδ2Vγ9 T cell proliferation (Lindberg et al., 2013) suggesting that phosphoantigens both are produced in and released from these cells. In a recent study we show that this occurs at all parasite blood stages and not only, as previously thought, when the erythrocytes rupture and release parasites (Liu et al., 2018).
Ingela Parmryd
Principal Investigator
Affiliation:
Department of Medical Biochemistry and Cell Biology,
Institute of Biomedicine
Group members
Jeremy Adler, PhD, Research Engineer
Sven-Göran Eriksson, BSc, Statistician
Chi-wen Huang, PhD student
Issac Thiria
Collaborators
Dr. Helgi Birgisson, Uppsala University Hospital, Sweden
Dr. Stefan Wennmalm, ALMI, SciLifeLab Stockholm, Sweden
Dr. Patrick Happel and Ms. Astrid Gesper, Ruhr-Universität Bochum, Germany
Prof. Benedikt Kost & Ms. Carolin Fritz, Universität Erlangen-Nürnberg, Germany
Prof. Gunilla Westermark, Uppsala University, Sweden