Genomic studies of neural tumors
Short description
Brain tumors are the most common form of solid tumors in children. Through genomic profiling of different neural tumor types, Frida Abel's research group's focus is to identify DNA variants that drive tumor progression, and genetic markers for risk stratification. Further functional analyses are performed to study the impact of DNA / RNA variants in tumor development. Genomic profiling is becoming one of the cornerstones of precision medicine in future healthcare. Our research enables implementation of new diagnostic methodology in clinical practice and contributes to increased understanding of neural tumor development. Genomic and transcriptomic profiling refines clinical diagnostics, facilitates risk stratification, and provides targets for precision treatment.
Genomic profiling of neural tumors – implementation into clinical diagnostics for precision care
Tumors of the nervous system (brain tumors and peripheral nervous system tumors) are the most common solid tumors in children, and unfortunately, many subtypes continue to have a suboptimal long-term outcome. Many patients suffer from long-term sequelae of surgery, cytotoxic chemotherapy, and radiotherapy. In a variety of cancer types, the identification of subtype specific tumor-driving mutations has been a critical initiating step in the path towards targeted therapy.
The growing need to simultaneously interrogate a large number of loci within a clinically relevant timeframe has directed a shift from pure histopathology towards genomic profiling by large-scale sequencing. Clinical genomics has the potential to refine tumor classification, stratify patients into treatment groups, identify druggable targets and malignancy markers, and allow treatment follow-up via tumor-specific biomarkers.
Our research aim is to 1) utilize new sequencing technologies for screening of novel genomic and transcriptomic events, 2) gain functional understanding of biological consequence of novel somatic and germline DNA alterations to inform targeted therapeutics, and 3) implement better molecular diagnostic methods and testing decision trees in clinical practice.
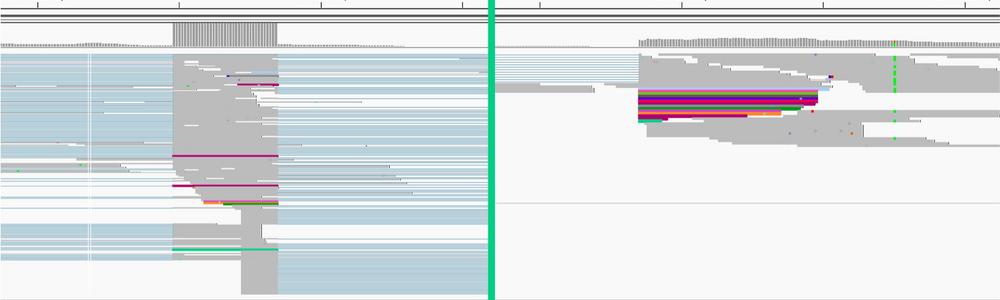
Tumor samples and corresponding blood has been collected since 2013 from all childhood brain tumor patients that have undergone surgery at Sahlgrenska. The new sequencing technologies are utilized to generate omics profiles of each patient’s RNA and DNA, forming the basis for molecular diagnostics and the search for druggable targets. By an integrative approach of different data levels and confirmational functional studies, the underlying mechanisms and core signaling pathways driving tumor progression are revealed.
Specific aims are:
(1) search for complex structural variants, e.g. gene fusions, promoter hijacking, inversions, translocations, deletions etc. through long-read sequencing of DNA and RNA in yet “unsolved” cases
(2) map germline and somatic tumor-driving DNA variants (single nucleotide variants, indels), structural variants, and copy-number changes through whole genome sequencing
(3) define core signaling pathways and prognostic markers by integrative analyses of different levels of omics data
(4) functionally validate the downstream effect of mutations and fusion genes.
The feasibility of this project is based on a strong network of collaborators spanning from the clinic (pediatric oncologists, neurosurgeons, neuropathologists) to advanced bioinformatics and experimental medicine. Our research increases the diagnostic yield, improves risk assessment and increases the chances of finding therapeutic targets for patients with neural tumors. Also, our research gains fundamental knowledge of tumor development and progression. Incorporation of genomic information into neuro-oncology diagnostics leads to improved outcomes and results in precision medicine for patients with nervous system (CNS and PNS) malignancies.

Frida Abel
Principal Investigator
Affiliation:
Department of Laboratory Medicine,
Institute of Biomedicine
Group members
Lily Deland (MSc, PhD student)
Angelica Ljungberg (MSc, assistant research scientist)
Eddie Vuong (MSc, engineer, assistant research scientist)
Simon Keane (external post doc collaborator)