Human genomics and bioinformatics
Short description
Erik Larsson Lekholm's research group is tackling a variety of problems in human computational genomics, with emphasis on basic cancer science. Additionally, some of our projects are more translational and involve collaboration with clinicians at the Sahlgrenska University Hospital. We also develop of bioinformatics tools to facilitate analysis of large-scale genomics data.
Massively parallel sequencing and other high-throughput methodologies enable detailed mapping of the genomic and transcriptomic changes that underlie the development of cancer and other diseases. Rapid accumulation of genetic information in the public domain makes it increasingly feasible to test biological hypotheses using computational methods and available data. We are tackling a variety of problems in human computational genomics, with emphasis on cancer.
A major focus of our efforts in cancer genomics is to better understand how somatic mutations arise and distribute across cancer genomes, and how this can be modelled computationally. This holds the promise of improving detection of signals of positive selection (i.e. potentially actionable driver alterations) in large-scale sequencing data from tumors.
Our research interests also include development of bioinformatics tools to facilitate analysis of large-scale genomics data, non-coding alterations in cancer, and mitochondrial DNA alterations.
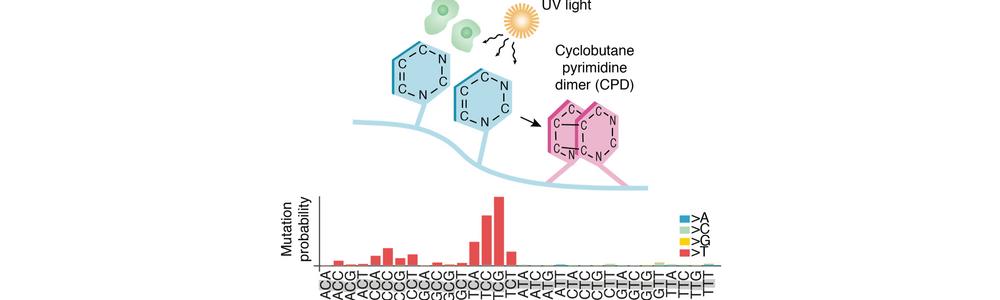
We have our own wet laboratory to complement our work in computational genomics. There, we are using high-throughput sequencing to map damage to DNA induced by ultraviolet light at high resolution, which has given important insights into mutational patterns in skin tumors.
Some of our projects are more translational and involve collaboration with clinicians at the Sahlgrenska University Hospital.

Erik Larsson Lekholm
Principal Investigator
Affiliation:
Department of Medical Biochemistry and Cell Biology,
Institute of Biomedicine

Group members
Alice Schiller, PhD student
Linnea Ögren, PhD student
Markus Lindberg, PhD, staff scientist
Kerryn Elliott, PhD, staff scientist
Isabella Muylaert, PhD, staff scientist